eLife 2022;11:e72483 DOI: 10.7554/eLife.72483
Rui Zheng, Yonglan Du, Xintai Wang, Tailin Liao, Zhe Zhang, Na Wang, Xiumao Li, Ying Shen, Lei Shi, Jianhong Luo
Department of Neurobiology and Department of Rehabilitation of the Children’s Hospital, Zhejiang University School of Medicine, China; NHC and CAMS Key Laboratory of Medical Neurobiology, Ministry of Education Frontier Science Center for Brain Research and Brain Machine Integration, School of Brain Science and Brain Medicine, Zhejiang University, China; 浙江大学
Department of Orthopedic Surgery, the Second Affiliated Hospital, Zhejiang University School of Medicine, China; Department of Physiology and Department of Neurology of First Affiliated Hospital, Zhejiang University School of Medicine, China;
JNU-HKUST Joint Laboratory for Neuroscience and Innovative Drug Research, Jinan University, China; Division of Life Science and The Brain and Intelligence Research Institute, The Hong Kong University of Science and Technology, China; Innovative Institute of Basic Medical Sciences of Zhejiang University (Yuhang), China
Abstract
Dynamic microtubules play a critical role in cell structure and function. In nervous system, microtubules are the major route for cargo protein trafficking and they specially extend into and out of synapses to regulate synaptic development and plasticity. However, the detailed depolymerization mechanism that regulates dynamic microtubules in synapses and dendrites is still unclear. In this study, we find that KIF2C, a dynamic microtubule depolymerization protein without known function in the nervous system, plays a pivotal role in the structural and functional plasticity of synapses and regulates cognitive function in mice. Through its microtubule depolymerization capability, KIF2C regulates microtubule dynamics in dendrites, and regulates microtubule invasion of spines in neurons in a neuronal activity-dependent manner. Using RNAi knockdown and conditional knockout approaches, we showed that KIF2C regulates spine morphology and synaptic membrane expression of AMPA receptors. Moreover, KIF2C deficiency leads to impaired excitatory transmission, long-term potentiation, and altered cognitive behaviors in mice. Collectively, our study explores a novel function of KIF2C in the nervous system and provides an important regulatory mechanism on how activity-dependent microtubule dynamic regulates synaptic plasticity and cognition behaviors.
動的な微小管は、細胞の構造と機能に重要な役割を果たしている。神経系では、微小管は荷電タンパク質の主要な輸送経路であり、シナプスの内外に特別に伸展してシナプスの発達と可塑性を制御している。しかし、シナプスや樹状突起におけるダイナミックな微小管の動きを制御する詳細な脱重合メカニズムは、いまだ不明であった。本研究では、神経系での機能が知られていない動的微小管解重合タンパク質であるKIF2Cが、シナプスの構造的・機能的可塑性に極めて重要な役割を果たし、マウスの認知機能を制御していることを明らかにした。KIF2Cは、その微小管脱重合能力により、樹状突起の微小管ダイナミクスを制御し、ニューロンの樹状突起スパインへの微小管侵入を神経細胞活動依存的に制御していることが知られている。RNAiノックダウンおよびコンディショナルノックアウト法を用いて、KIF2Cがスパインの形態およびAMPA受容体のシナプス膜発現を制御していることを明らかにした。さらに、KIF2Cの欠損は、興奮性伝達、長期増強の障害や、マウスの認知行動の変化をもたらすことを明らかにした。本研究は、神経系におけるKIF2Cの新たな機能を明らかにするとともに、活動量に依存した微小管の動態がシナプス可塑性や認知行動を制御する重要な制御機構を提供するものである。
Editor’s evaluation
In this manuscript, the authors report a set of exciting and novel data suggesting a critical role of the microtubule depolymerizing kinesin-13 KIF2C/MCAK in glutamate receptor trafficking, synaptic plasticity and behaviours related to learning and memory. The authors use multiple experimental approaches, ranging from electron microscopy to mouse behavioral analysis to support the conclusions, which are convincing and provide novel insights into the in vivo functions of KIF2C in the regulation of excitatory synaptic structure and function and cognitive behaviors.
本論文では、微小管伸長因子KIF2C/MCAKがグルタミン酸受容体の輸送、シナプス可塑性、学習・記憶に関する行動に重要な役割を果たすことを示唆する一連の興味深い新規データを報告している。著者らは、電子顕微鏡法からマウスの行動分析に至るまで、複数の実験的アプローチを用いて結論を支持しており、その結論は説得力があり、興奮性シナプスの構造・機能および認知行動の制御におけるKIF2Cの生体内での機能に対する新しい洞察を与えるものであった。
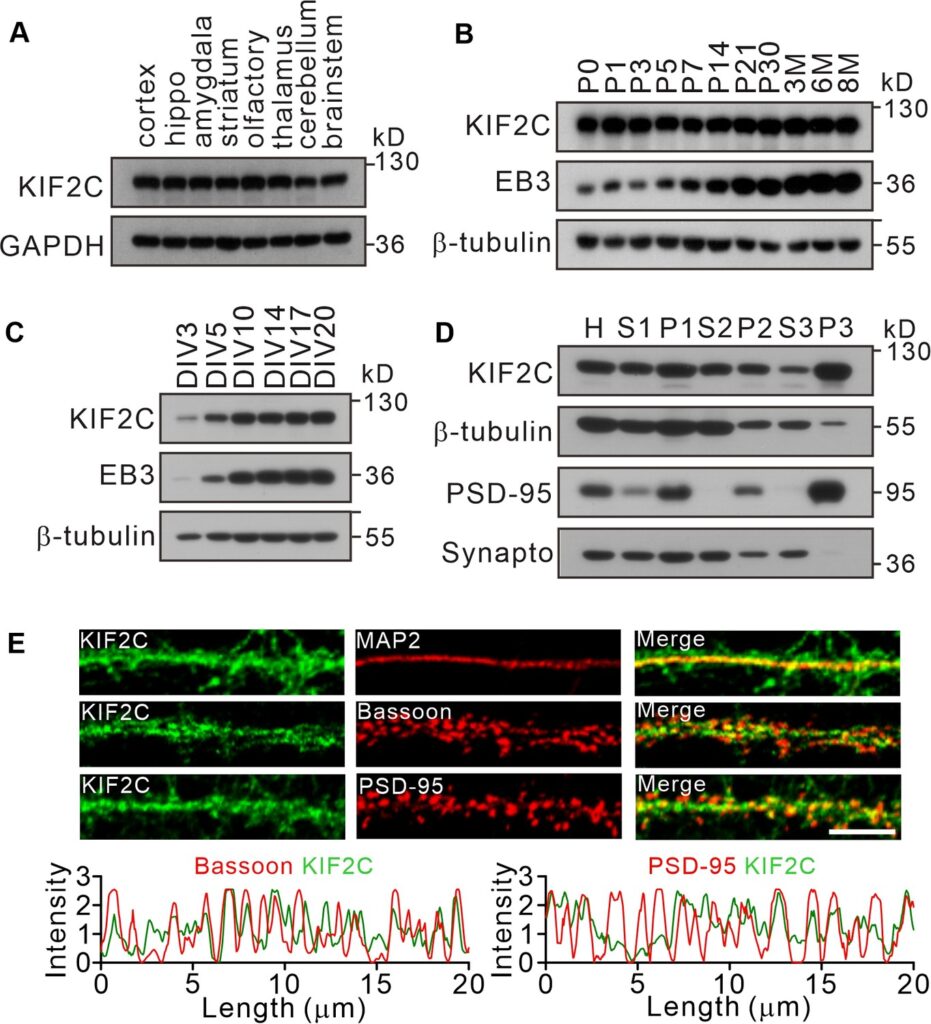
(A) Expression of KIF2C in different mouse brain regions by immunoblotting. (B) Expression of KIF2C and EB3 in mouse hippocampus at different day in vitro. (C) Developmental expression patterns of KIF2C and EB3 in cultured hippocampal neurons. (D) Subcellular distribution of KIF2C in mouse hippocampus. H, homogenate; S1, low-speed supernatant; P1, nuclei; S2, microsomal fraction; P2, synaptosomal fraction; S3, presynaptosomal fraction; P3, postsynaptic density and synapto, synaptophysin. (E) Representative images of hippocampal neurons double-labeled with antibodies against KIF2C, PSD-95 (postsynaptic marker) and Bassoon (presynaptic marker). Scale bar, 5 μm.
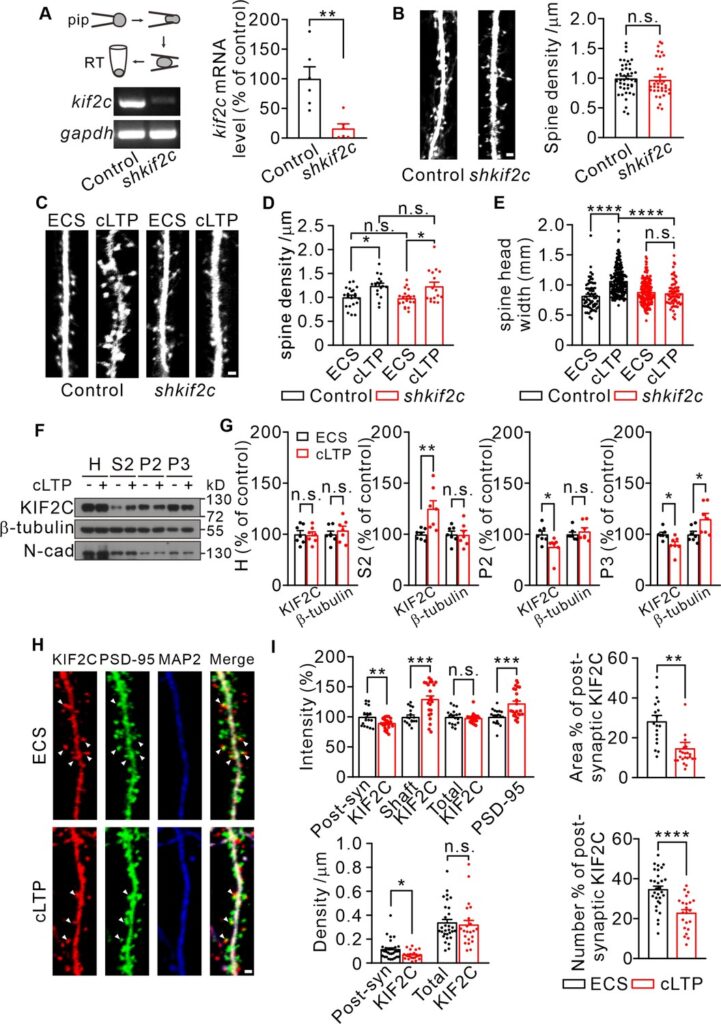
(A) Electrophoresis of kif2c and gapdh amplicons from individual control and shkif2c hippocampal neurons. Histograms show percentage changes of kif2c mRNA levels (% of control, **p < 0.01, n = 6, Student’s t-test). (B) Cultured hippocampal neurons were infected by control shRNA and shkif2c lentivirus at DIV 7–10, and captured at DIV 18–20. GFP was used to label neuron volume. Scale bar, 1 μm. Spine number per 1 μm (right), n = 20 neurons from three independent culture. Student’s t-test. (C–E) Cultured hippocampal neurons were infected by control shRNA and shkif2c lentivirus at DIV 7–10, and incubated with ECS or glycine (200 μM) for 10 min at DIV 18–20. Scale bar, 1 μm. Spine density (D) and spine head width (E) are calculated. Spine density, n = 17 neurons from three independent culture. Spine head width, n = 24 neurons from three independent culture. n.s., p > 0.05, *p < 0.05, ****p < 0.0001; One way-ANOVA for spine density and head width (post hoc comparison).(F) Subcellular fraction of KIF2C in cultured hippocampus neurons after 10 min of glycine treatment. N-cadherin (N-cad) was served as negative control. H, Homogenate; S2, microsomal fraction; P2, synaptosomal fraction and P3, postsynaptic density from ECS and cLTP treated neurons. (G) Quantification of KIF2C and β-tubulin accumulation with or without cLTP treatment in each fraction. *p < 0.05, **p < 0.01; Student’s t-test, n = 7 experimental repeats. (H) Representative images of co-localization of KIF2C (red), PSD-95 (green), and Microtubule associated protein 2 (MAP2) (blue). Scale bar, 1 μm. (I) The percentage of KIF2C puncta that were colocalized with PSD-95 puncta decreased after cLTP treatment. n = 20 neurons from three cultures. n.s., p > 0.05, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; Student’s t-test.
